 
Head(datlm) # block grp x1 x2 resp predlm fit lwr upr These columns can be bound to dat for plotting. predslm = predict(fitlm, interval = "confidence") I used the default and so get a 95% confidence interval for each predicted value. □Īdding interval = "confidence" returns a three column matrix, where fit contains the fitted values and lwr and upr contain the lower and upper confidence interval limits of the predicted values, respectively. I’m skipping the assumption-checking step here. What about confidence intervals? The predict() function for lm objects has an interval argument that returns confidence or prediction intervals, which are appropriate to use if model assumptions have been reasonably met. For example, ?predict.lme will take you to the documentation for the predict() function for lme objects fit with nlme::lme(). You can go to the help page for the predict() function for a specific model type. # see '?methods' for accessing help and source code (Also see, e.g., methods(class = "lm") for functions available for a specific model type.) methods("predict") # predict.ar* predict.Arima* Since I’ve already loaded package nlme you can see predict.lme and predict.gls along with many others. For example, methods("predict") lists all the different model objects that have specific predict() functions. You can check if the model you are using has a predict function via methods(). For many model types the predictions can be extracted from the fitted model via the predict() function. These predicted values can then be used for drawing the fitted line(s). We can instead fit a model and extract the predicted values. To free ourselves of the constraints of geom_smooth(), we can take a different plotting approach. If you want parallel lines instead of separate slopes per group, geom_smooth() isn’t going to work for you. In the plots above you can see that the slopes vary by grp category. 15, aes(fill = grp))Įxtracting predicted values with predict() ggplot(dat, aes(x = x1, y = resp, color = grp) ) + I increased the transparency of the ribbons by decreasing alpha, as well, since adding confidence ribbons for many fitted lines in one plot can end up looking pretty messy. I used fill to make the ribbons the same color as the lines. Here is the same plot with a 95% confidence envelope (the default interval size) as a ribbon around the fitted lines. This is a linear model fit, so I use method = "lm". Confidence intervals can be suppressed using se = FALSE, which I use below. 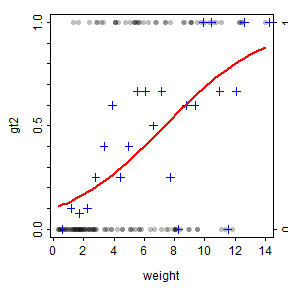 
This can be great if you are plotting the results after you’ve checked all assumptions but is not-so-great if you are exploring the data. 
I’m going to plot fitted regression lines of resp vs x1 for each grp category.īy default you will get confidence intervals plotted in geom_smooth(). Supported model types include models fit with lm(), glm(), nls(), and mgcv::gam().įitted lines can vary by groups if a factor variable is mapped to an aesthetic like color or group. The geom_smooth() function in ggplot2 can plot fitted lines from models with a simple structure. Plotting separate slopes with geom_smooth() These data are from a blocked design, and the block variable is available to be used as a random effect. This dataset has one response variable, resp, along with two continuous ( x1, x2) and one categorical ( grp) explanatory variables. I created a dataset to use for fitting models and used dput() to copy and paste it here. I’m going to set the ggplot2 theme to theme_bw(). First I’ll load the packages I’m using today. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
